FMD Modules Manual
The Forensic Microbiome Database (FMD) is a human microbiome analysis resource that correlates publicly available 16S rRNA sequence data obtained from multiple body sites to metadata as it relates to forensics, using several analytic techniques:
- Looking at the taxonomic distribution of individual samples from the collected publicly available 16S rRNA sequence data;
- Comparing the taxonomic distribution of multiple samples from the collected publicly available 16S rRNA sequence data;
- Locating a user-supplied sample by comparing its taxonomic distribution to all the publicly available 16S rRNA sequence data;
Home Page
The FMD homepage allows users to connect microbiome sequences to geography. The users can explore the publicly available data, use the available tools, and load their own data to use the tools. The home page can be accessed from any other page by clicking on the icon in the top left, on the home in the header menu or in the footer menu. Users can access a more personalized FMD by logging on by selecting the Sign In (1) button on the top right or by Signing Up (1) if the user has not already.
Header Menu
The header menu allows to explore the database, access the tools, and load their own data. The menu includes:
- Home: goes to the home page
- Analysis: Provides the tools to explore the connection between geography and microbiota
- Geographic Location Prediction: Predict where a sample is from based on similarity to samples in the database
- Taxonomic Distribution Viewer: Examine the taxonomic composition of database samples using interactive viewers
- Comparative Taxonomic Distribution Viewer: Compare taxonomic compositions of multiple database samples
- Data: Allows the user to examine and upload data, both their own and the sites
- Data statistics: Summary of the data available in the FMD
- Download Data: Coming Soon!
- Submit Data: Coming Soon!
- Support: Provides help and information for the user to use the site
- User Manual
- Standard Operating Procedures: Coming Soon!
- Frequently Asked Questions
- Resource Policy
- Cite FMD
- Contact

Body
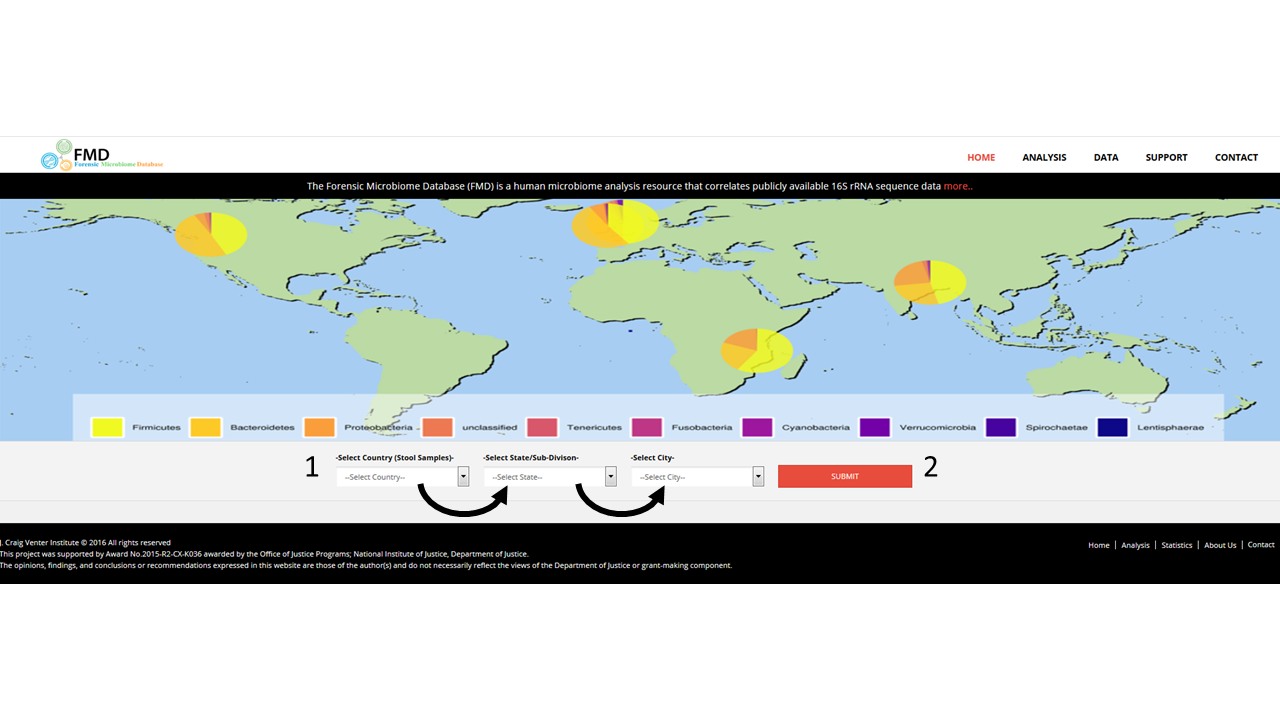
The body of the main page contains a map showing the proportion of stool samples from different countries are from different Phyla. Underneath the map is a set of menus (1) that allow users to create an interactive Krona pie chart of the distribution of the genera from different stool samples from different cities. The menus are auto-filled after the menu to the left is selected, and the Krona interactive image is loaded by clicking on the submit button (2)..
Footer
The footer contains several additional links, including:- JCVI: this website is supported by JCVI and the DOJ under the grant cited below
- Home: Goes to the home page
- Prediction: Predict where a sample is from based on similarity to samples in the database
- Statistics: Summary of the data available in the FMD
- About Us: Gives a brief description of the FMD project
- Contact: provides a form to contact us

Predict a Geographic Location
The FMD allows users to estimate the geographic origin of a sample or group of samples by comparing its 16SrRNA community profiles to the samples collected worldwide by the FMD project. To access this tool, mouse over the Analysis menu option (1) and select "Geographic Location Prediction"(2).
Loading Files
Once on the page, FMD prompts users for two files that describe the taxonomic distribution of the sample to assay. The two files are outputs from the mothur open-source microbial community analysis pipeline: the shared (file.shared) file- a file containing the counts of each operational taxonomic unit (OTU) across samples- and the taxonomic (file.taxonomy) file- a file containing the taxonomic ID hit for every OTU. Acceptable file names are limited to "otu.shared" for the shared file and "otu.taxonomy" for the taxonomy file, all in lower case. To run the geolocation tool, load in the two files, "otu.shared" (1) and "otu.taxonomy" (2) and select "Submit For Prediction" (3) to load the results page.
File Formats
Examples of the two files are available on the page. The shared file contains the raw counts of each OTUs across samples. The header contains the names of each OTU labeled as "otu1, otu2, otu3... otuN" (1) and the second row contains the name, the total count of identified OTUs and the subsequent columns show the counts for each OTU across the sample (2).
The taxonomy file contains matches between the OTUs in the shared file and its taxonomic classification. The header is required to be "OTU\tTaxonomy\n" (1);
Each OTU is defined as a group of sequences sharing >97% 16S rDNA similarity.
Results Page
The results page shows the longitude and latitude of the sample from the publicly available 16S rRNA data that best matches the inputted sample using Google Maps (1). The distance or degree of similarity between uploaded sample and reference data is calculated using the Sorensen Similarity Index (2). The Similarity Index uses the OTU presence and abundance counts between two samples and goes from 1 (=100% matching data) to 0 (=0% matching data).
16S rRNA-based Taxonomic Distribution for a Geographic Location
The FMD allows users to examine the taxonomic distribution of all the samples in the FMD collected in a single geographic location from a single body site. It can be accessed from the main page in two ways, via the menu, selecting Taxonomic Distribution viewer under Analysis (1) or using the menus under the map (2). A country (the left-most menu) has to be selected first. The state menu is then auto-populated with available data from the selected country and so on. A country, state and city all have to be selected. After each have been chosen, click on the submit button (3).
Selecting Body Sites and Geographic Locations
At the Taxonomic Distribution Viewer page, the FMD data can be initially searched by either body site or by geography (country) (1). It is required to select one before continuing by clicking Next (2).
On the selection page, each menu is auto-populated after the selection on the right is made (1). To view the image showing the distribution of the taxonomic assignment, click on the submit button (2). Both the body site and the country are required to be selected to view the image. To return to the initial selection step, click the step 1 button (3).
Result Page
The taxonomic image is shown as a Krona pie chart, where the percentage of sequenced 16S rRNAs assigned to each taxonomic unit is shown as a percentage of the circumference (1). Different taxonomic levels are shown along the radius of the chart (2). Taxonomic regions in the graph can be expanded by double clicking on the colored areas. Geographic sub-divisions of the selected geographic location can be accessed using the menu on the top left (3). Further information on Krona pie charts and their functionality can be found using this link or by clicking on the "?" button on the graph (4).
Comparative Taxonomic Distribution Viewer
The FMD allows users to compare the taxonomic distribution of multiple geographic locations from the samples in the FMD from a single body site. To access it, select the "Comparative Taxonomic Distribution Viewer" (1) option under the Analysis Menu (2).
Selecting Initial Body Sites and Geographic Locations
At the Comparative Taxonomic Distribution Viewer page, the FMD data can be initially searched by either body site or by geography (country) (1). It is required to select one before continuing by clicking next (2). Also, the taxonomic level for the analysis can be selected (3).
On the initial selection page, each menu is auto-populated after the selection on the right is made (1). To view the image showing the distribution of the Taxonomic assignment, click on the submit button (2). Both the body site and the country are required to be selected to view the image. To return to the initial selection step, click the step 1 button (3).
Selecting Other Body Sites and Geographic Locations
On the secondary selection page, all additional samples with the same body site and geographic level are shown (1). To select a geographic location to be compared to the original, select one of the choices in the left box (1) and choose Add (2). It will then appear in the right box (3). To view the comparison between the initial selected location and all the locations in the right box, select submit.
Result Page
The results page shows the proportion of the top twenty groups at the selected taxonomic levels in each of the selected geographic locations as a bar graph (1), using the ranks from the initially selected location (2). Hovering over the bars will show the exact percentage as a text box. The image can be saved in multiple formats (3).
FMD Data Statistics
The data statistics page contains several descriptive counts of the data available in the FMD. These include:
- The number of high quality samples per body site
- The number of these samples in each body site in each geographic location, including country, state and city
- The number of these samples in each geographic location, including country, state and city from all body sites
- The number of these samples in each geographic location, including country, state and city broken down by study
